PSA Laboratory Software, Michigan State University Protein Structural Analysis Laboratory. Download digital pipe fitter keygen download pc. MSU SLIDE ('Screening for Ligands by Induced-fit Docking, Efficiently') represents a. You can download a free copy of Acrobat Reader from here. Molecular Docking Freeware Software Molegro Virtual Docker for Mac OS v.4.2 Handles all aspects of the molecular docking process from preparation of the molecules to determination of the potential binding sites of the target protein, and prediction of the binding modes of the ligands. Molecular Docking. DOCK University of California San Francisco. AutoDock Scripps Research Institute. Molegro Virtual Docker Molegro ApS, University of Aarhus, Denmark. Hex Protein Docking. University of Aberdeen, UK. GRAMM Protein docking software Center for Bioinformatics, University of Kansas, USA. Protein-protein docking software tools| Interaction data analysis. ATTRACT is a program available through a desktop version and a web application. 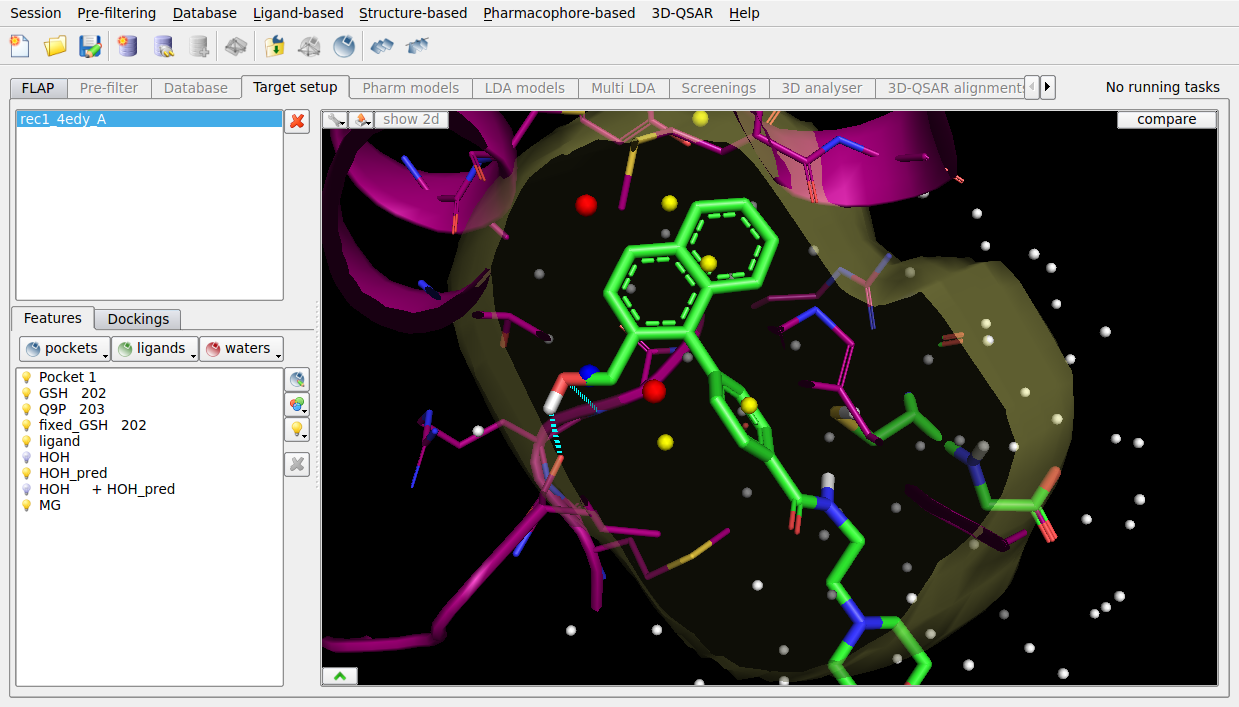
• • • • • • • • • • Ruth Huey William Lindstrom Scott Halliday AutoDock is a suite of automated docking tools. It is designed to predict how small molecules, such as substrates or drug candidates, bind to a receptor of known 3D structure. Current distributions of AutoDock consist of two generations of software: AutoDock 4. 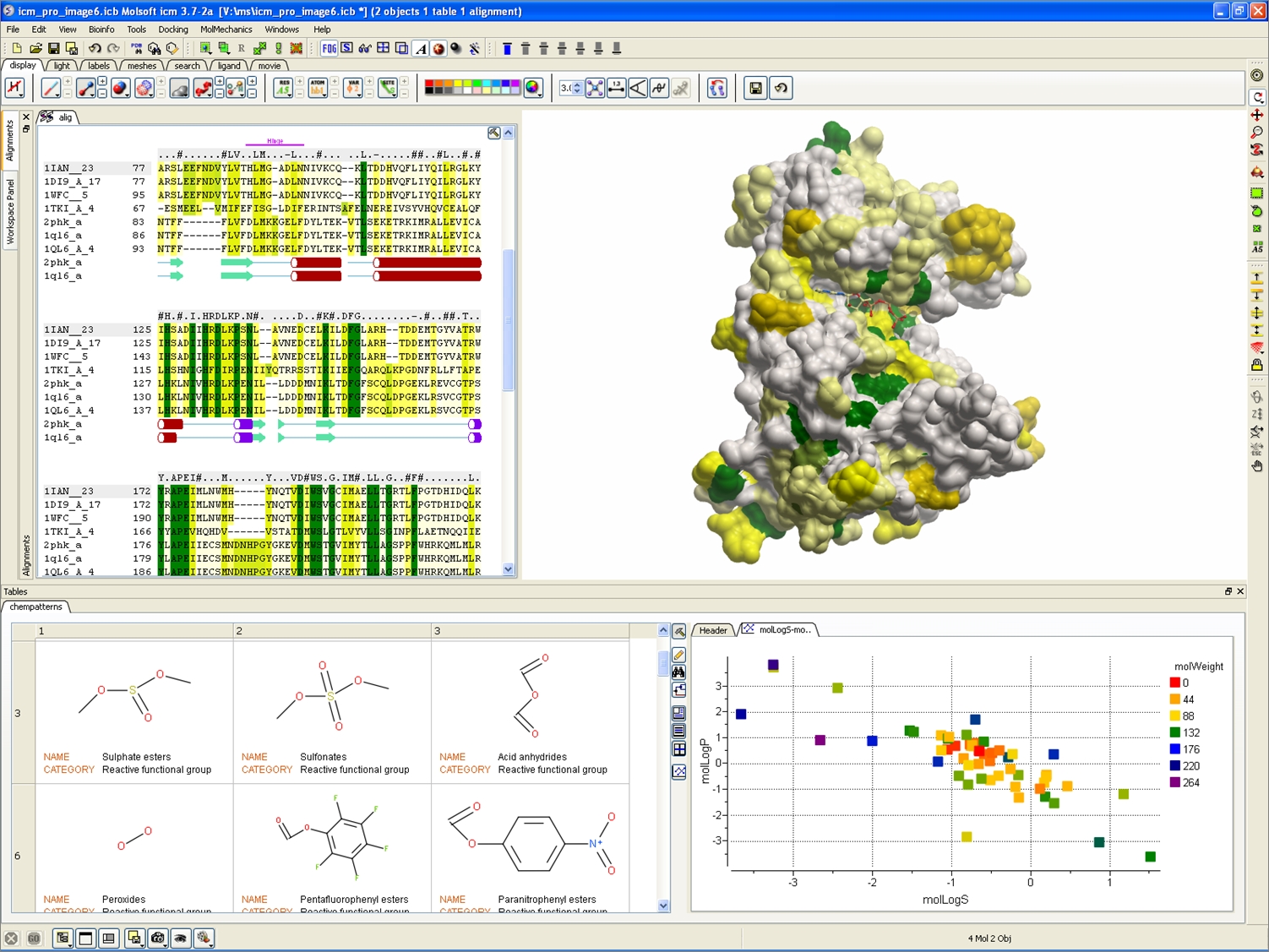 
AutoDock 4 actually consists of two main programs: autodock performs the docking of the ligand to a set of grids describing the target protein; autogrid pre-calculates these grids. In addition to using them for docking, the atomic affinity grids can be visualised. This can help, for example, to guide organic synthetic chemists design better binders. Does not require choosing atom types and pre-calculating grid maps for them. Molecular Docking SoftwareInstead, it calculates the grids internally, for the atom types that are needed, and it does this virtually instantly. We have also developed a graphical user interface called, or ADT for short, which amongst other things helps to set up which bonds will treated as rotatable in the ligand and to analyze dockings. AutoDock has applications in: • X-ray crystallography; • structure-based drug design; • lead optimization; • virtual screening (HTS); • combinatorial library design; • protein-protein docking; • chemical mechanism studies. AutoDock 4 is free and is available under the GNU General Public License. AutoDock Vina is available under the Apache license, allowing commercial and non-commercial use and redistribution. Click on the ' tab. And Happy Docking! What is AutoDock Vina? AutoDock Vina is a new generation of docking software from the Molecular Graphics Lab. It achieves significant improvements in the average accuracy of the binding mode predictions, while also being up to two orders of magnitude faster than AutoDock 4. Because the scoring functions used by AutoDock 4 and AutoDock Vina are different and inexact, on any given problem, either program may provide a better result. Detailed information can be found on the. Realtek 8812au wireless lan 802.11ac usb nic windows 10 driver. Docking Protein Software Free DownloadsJune 1, 2009 AutoDock 4.2 is faster than earlier versions, and it allows sidechains in the macromolecule to be flexible. As before, rigid docking is blindingly fast, and high-quality flexible docking can be done in around a minute. Up to 40,000 rigid dockings can be done in a day on one cpu. AutoDock 4.2 now has a free-energy scoring function that is based on a linear regression analysis, the AMBER force field, and an even larger set of diverse protein-ligand complexes with known inhibition constants than we used in AutoDock 3.0. The best model was cross-validated with a separate set of HIV-1 protease complexes, and confirmed that the standard error is around 2.5 kcal/mol. This is enough to discriminate between leads with milli-, micro- and nano-molar inhibition constants. Docking Protein Software Free SoftwareYou can read more about the new features in AutoDock 4.2 and how to use them in the. May 7, 2007 The introduction of AutoDock 4 comprises three major improvements: • The docking results are more accurate and reliable. • It can optionally model flexibility in the target macromolecule. • It enables AutoDock's use in evaluating protein-protein interactions. AutoDock 4.0 not only is it faster than earlier versions, it allows sidechains in the macromolecule to be flexible. As before, rigid docking is blindingly fast, and high-quality flexible docking can be done in around a minute. Up to 40,000 rigid dockings can be done in a day on one cpu. Free Molecular Docking SoftwareAutoDock 4.0 now has a free-energy scoring function that is based on a linear regression analysis, the AMBER force field, and an even larger set of diverse protein-ligand complexes with known inhibiton constants than we used in AutoDock 3.0. The best model was cross-validated with a separate set of HIV-1 protease complexes, and confirmed that the standard error is around 2.5 kcal/mol. This is enough to discriminate between leads with milli-, micro- and nano-molar inhibition constants. You can read more details about the new features in. AutoDock 4.0 can be compiled to take advantiage of new search methods from the optimization library, ACRO, developed by William E.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
